You're testing one or two diseases in animal models. That's like betting everything on red at the casino.
Sparx is a human-genetic target validation platform that uses natural genetic experiments to identify which of 2,000+ diseases has the strongest causal evidence for your molecular target—so you don’t commit to expensive preclinical validation that fail to translate to humans 90% of the time.

Animal models do not fully capture the complexity of the human disease. What works in mice does not predict well what works in humans.

Observational studies aren't better. They show correlation, not causation. Is your target causing the disease, or is the disease affecting your target?
By the time you discover you chose the wrong target-disease pair in Phase 2 trial, you've already invested millions and years of work.


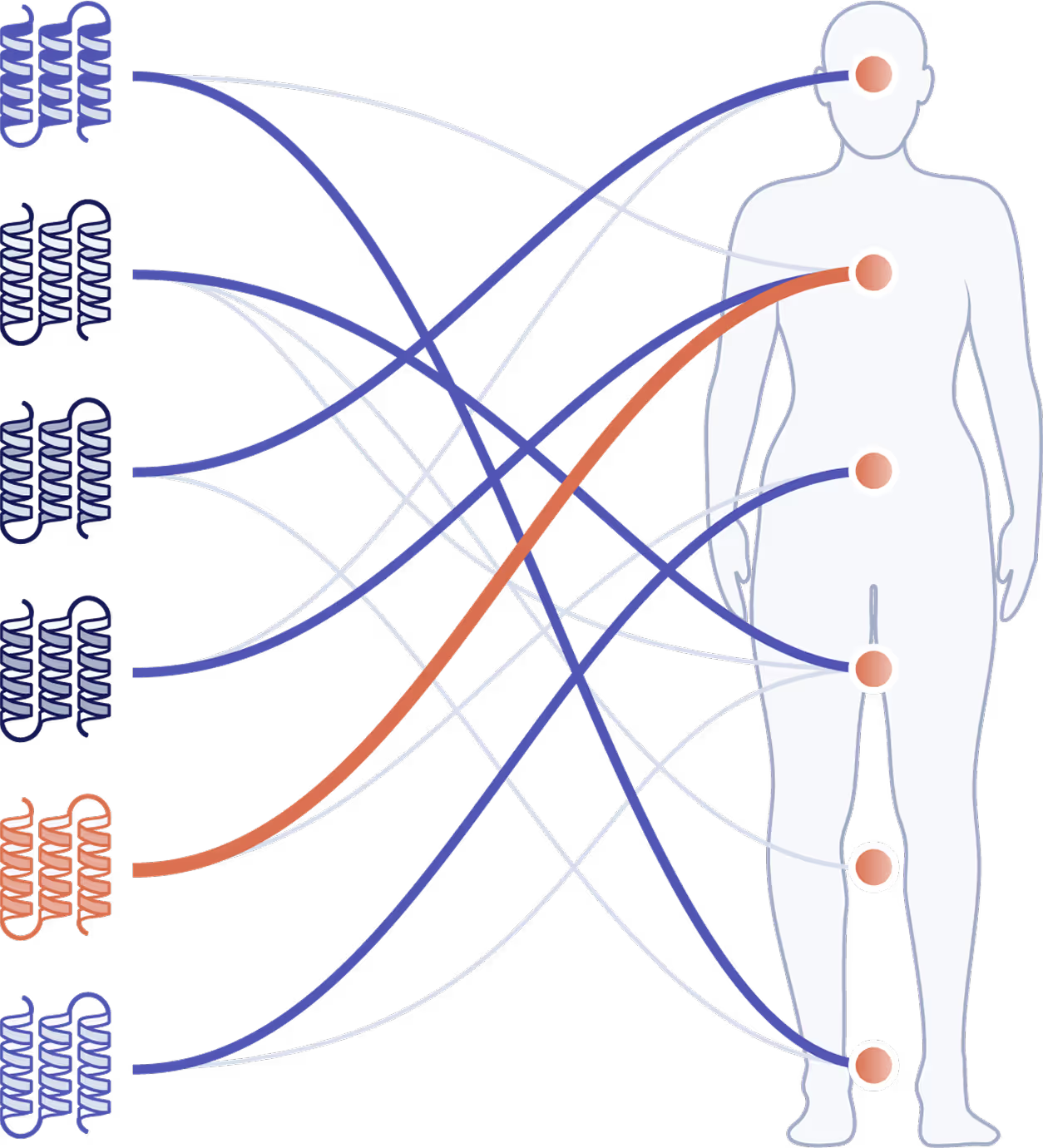
Test a single target across 2,000+ diseases and biomarkers to select the indication most likely to succeed—not just the first one you try.

Identify biomarkers of early efficacy to inform go/no go decisions in phase I clinical trials.
Prioritize the tissue of activity for a given target-disease pair.

Assess how a novel drug target performs, both in efficacy and safety, compared with established drug targets.
We built RETIN™ (Ranking Engine for Target–INdication pairs), a state-of-the-art algorithm that helps biotechs develop safer, more effective drugs for chronic diseases, by ranking molecular targets by their predicted efficacy and safety. RETIN™ is trained on large-scale human genetic data and validated using approved drug outcomes.
Tell us your molecular targets and share preclinical data worth considering. No wet lab work required. No IP transfer.
We use genetic validation from millions of humans to predict which target-disease pairs will succeed in clinic—before you file your IND.
Pursue your drug development program with confidence backed by human genetic data.
Short answer: no. Rare disorders already have established genetic targets. We focus on chronic diseases where finding the best target is challenging.
True, but successful therapies still target one or two proteins. We are finding targets that are central to the disease pathophysiology.
Yes. We use genetic causality that applies to any target-disease pair, not historical training data. Genetic variants exist for most human proteins.
No. To prevent hallucinations and availability bias, we don't use large language models. We analyze actual genetic data from millions of individuals to establish causal inference. We're asking: do people born with this genetic variant have different disease rates? That's causation, not correlation.